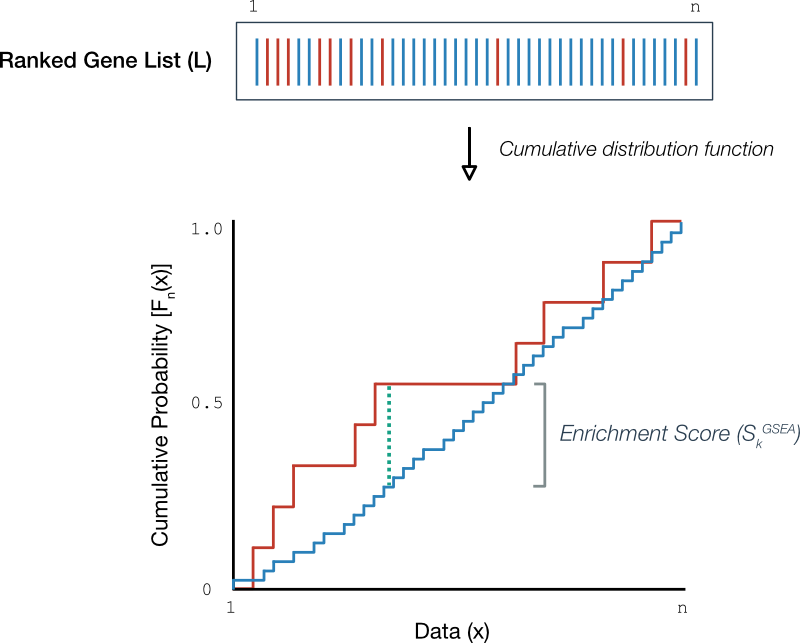
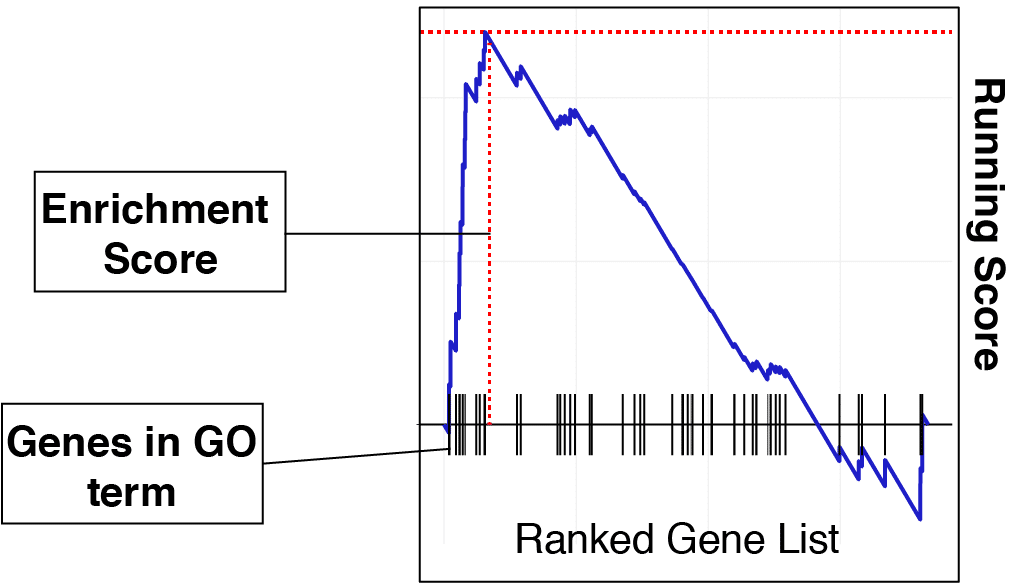
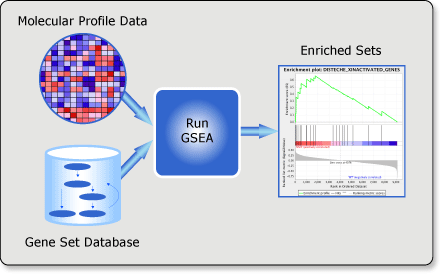
Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles | PNAS
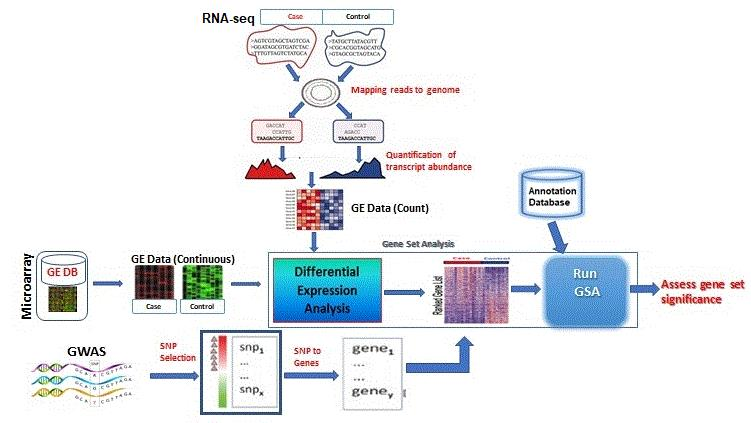
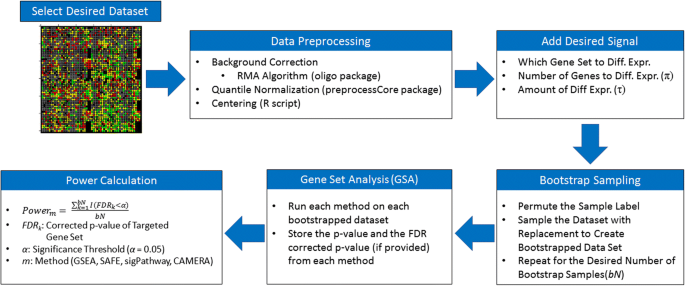
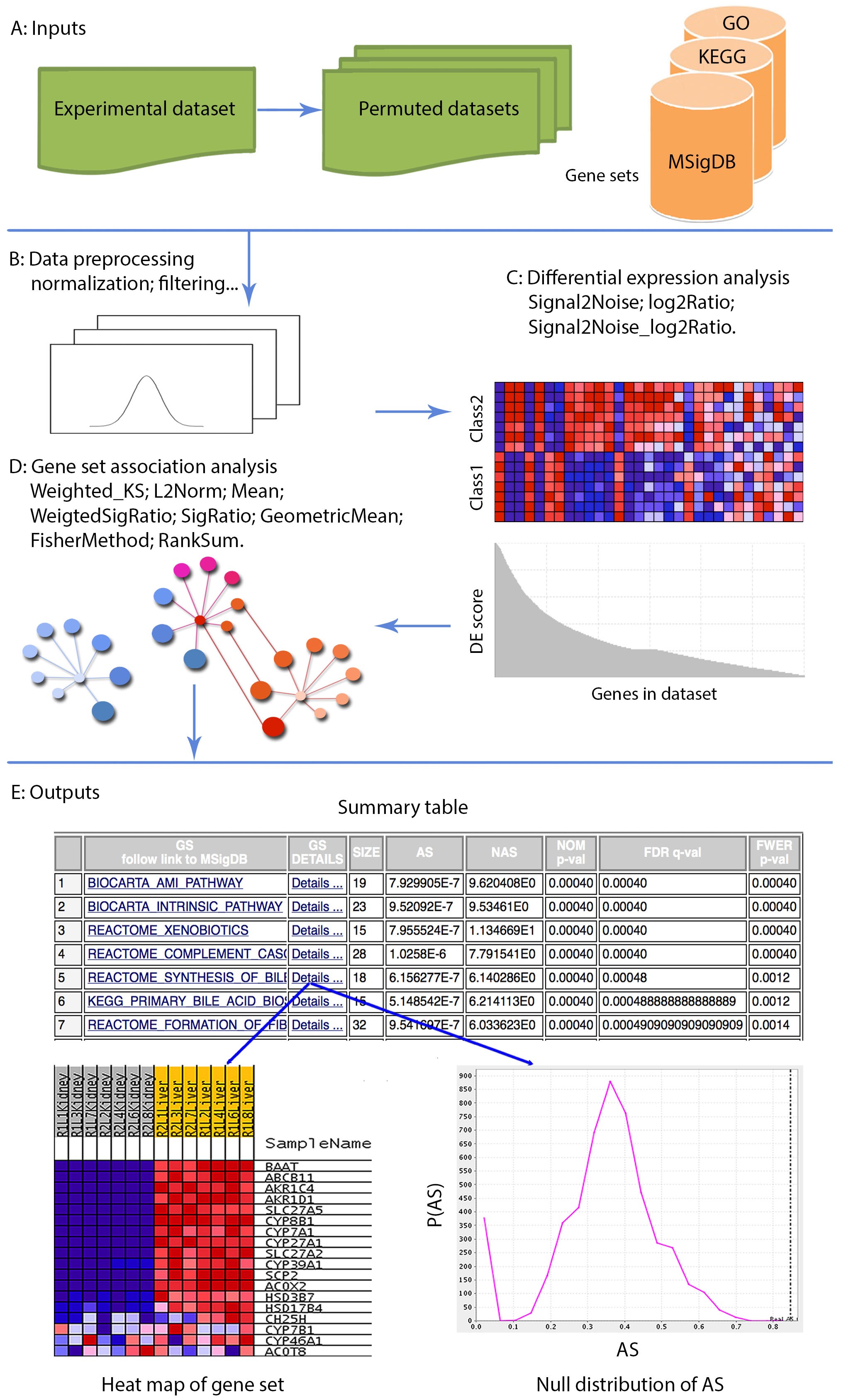
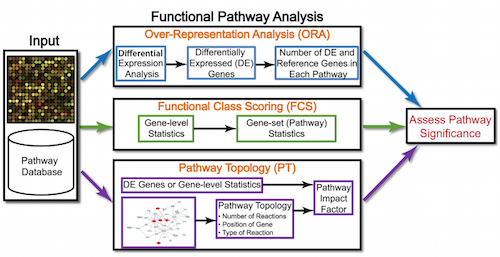
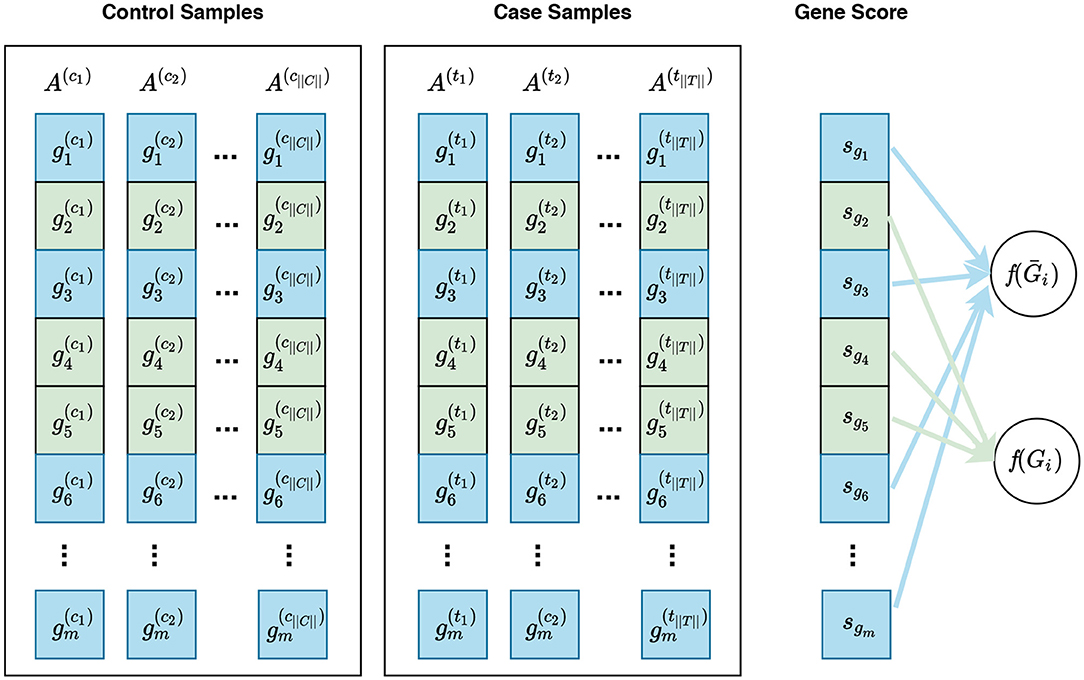
Entropy | Free Full-Text | Fifteen Years of Gene Set Analysis for High-Throughput Genomic Data: A Review of Statistical Approaches and Future Challenges
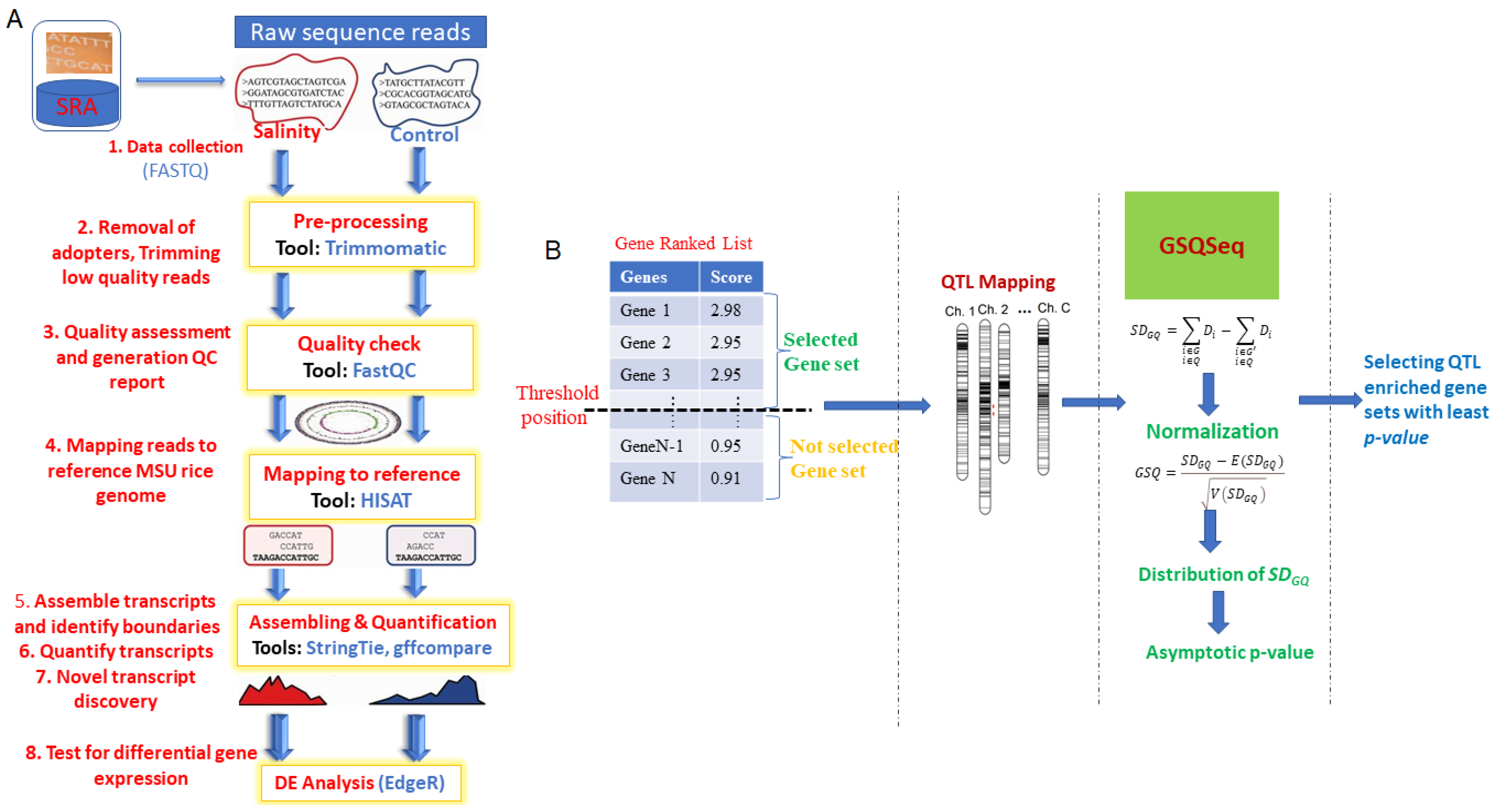
Entropy | Free Full-Text | Statistical Approach of Gene Set Analysis with Quantitative Trait Loci for Crop Gene Expression Studies

Signature-scoring methods developed for bulk samples are not adequate for cancer single-cell RNA sequencing data | eLife
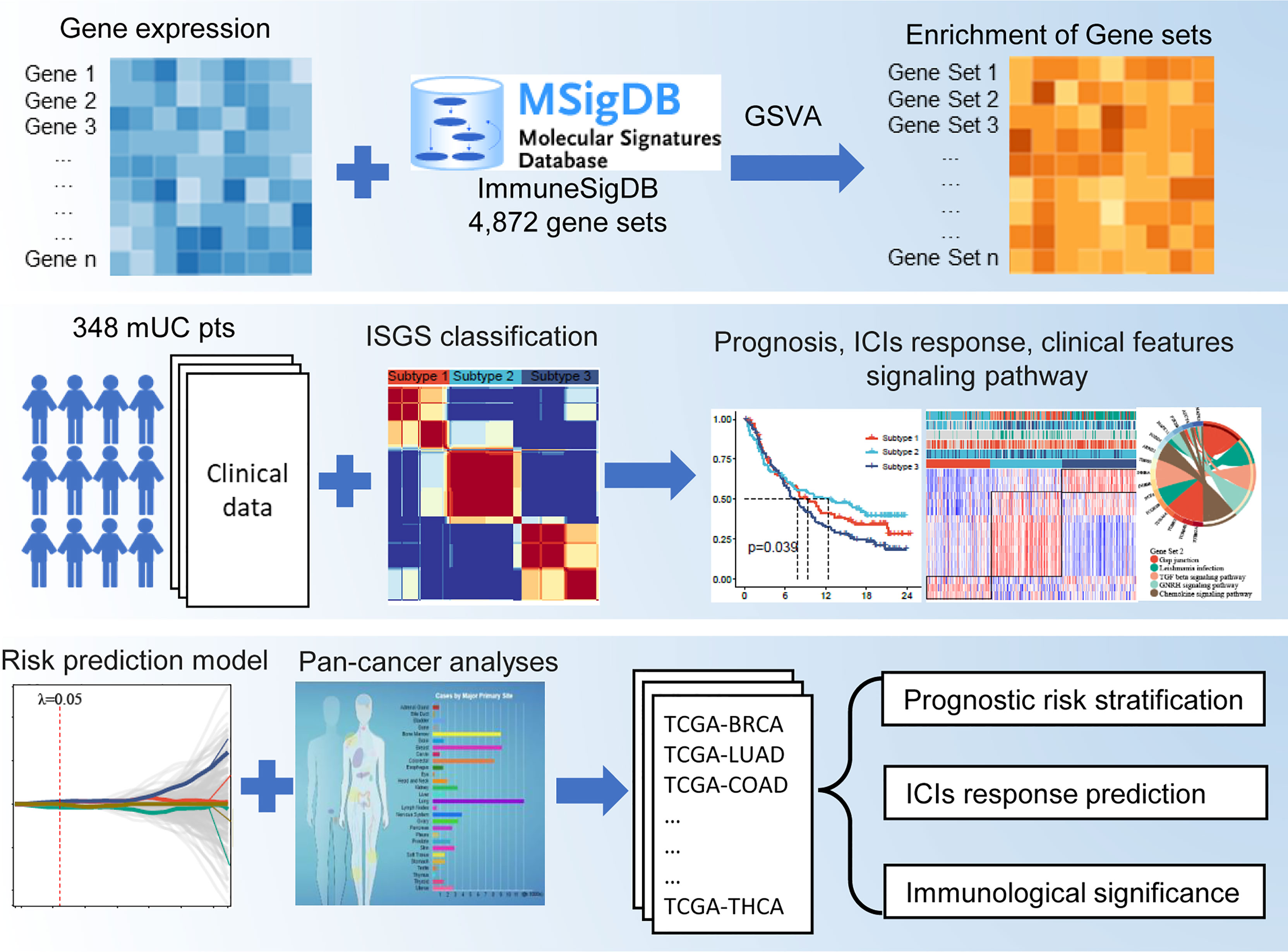
Frontiers | Immunologic Gene Sets Reveal Features of the Tumor Immune Microenvironment and Predict Prognosis and Immunotherapy Response: A Pan-Cancer Analysis
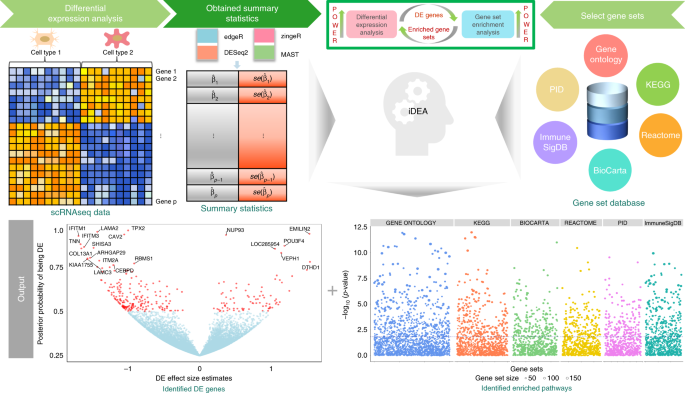
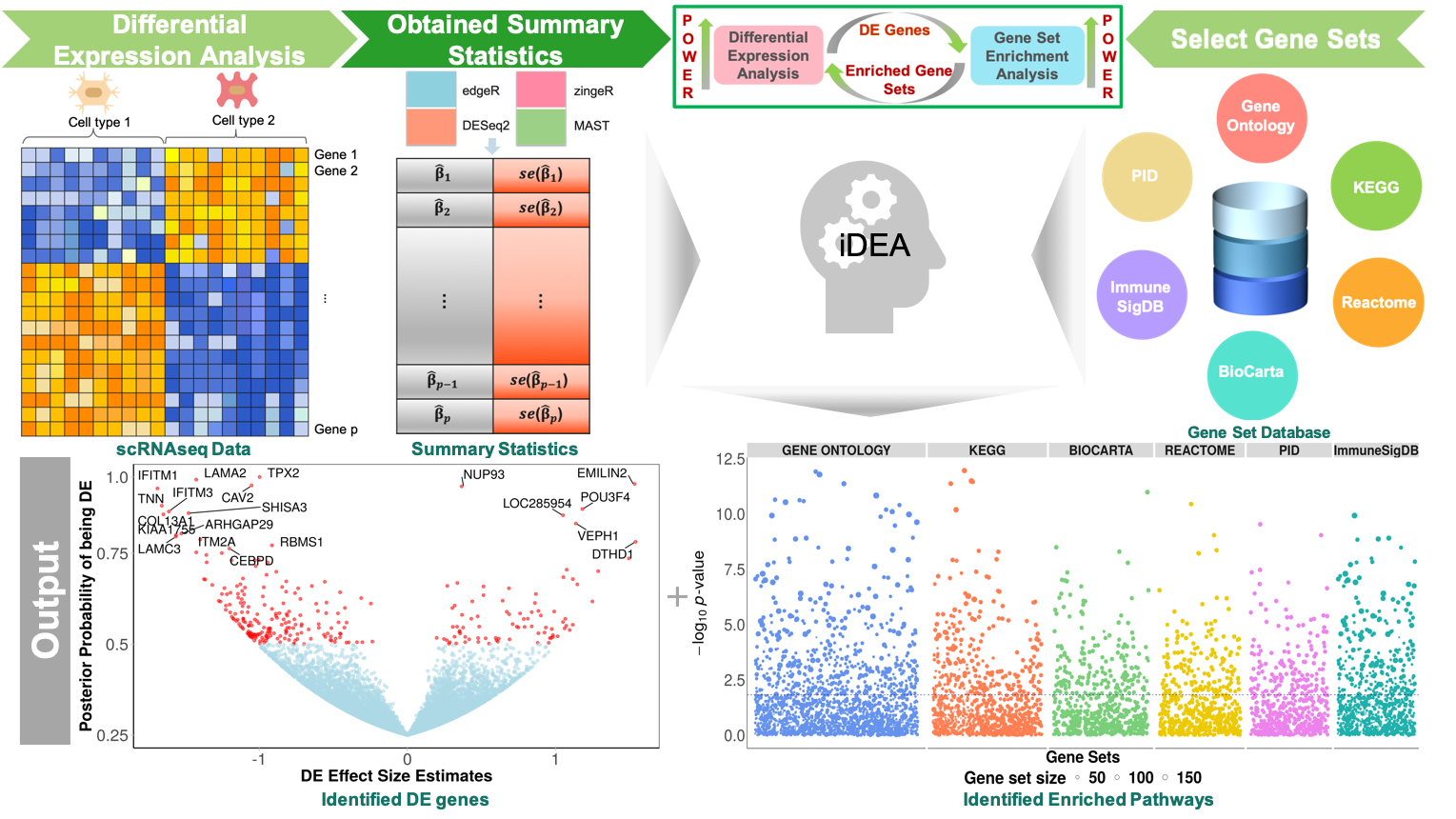
Integrative differential expression and gene set enrichment analysis using summary statistics for scRNA-seq studies | Nature Communications

Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles | PNAS